Diagnosis
-
MALDI-TOF in diagnostics
Establishment of MALDI-TOF in diagnostics of implant-associated or periprosthetic joint infections.
Oliver Bader, University of Göttingen, Germany
The current diagnostic procedure for the determination of infectious agents in implant-associated infections is laborious and slow. Such diagnostic specimens usually only have a low microbial density, with high concomitant cellular contamination, so that direct identification of living microbes without a culture step is not feasible. This culture step delays the diagnosis by at least one day, often much longer.
MALDInfect evaluates matrix-assisted laser desorption/ionization-time of flight (MALDI-TOF) mass spectrometry (MS)-based pathogen identification directly from diagnostic specimens (joints, explants, or tissues), to speed up diagnosis. The project is a collaborative effort between the Department of Trauma Surgery, Orthopedics and Plastic Surgery and the Institute for Medical Microbiology and Virology at the University Medical Center Göttingen, Germany. This interdisciplinary team allows us to bridge the whole diagnostic path from the patient to the laboratory and back.
The working hypothesis is that a clinical specimen can already be considered a suitable form of microbial culture, at least for monomicrobial growth. MALDI-TOF MS identification of microbes can be performed with protein fingerprints obtained from minute amounts of samples, down to approximately 105 cells. The main remaining challenge in MALDInfect is to develop proper (non-culture-based) enrichment and purification processes. In the course of the project, such protocols are being developed for three specimen groups, gradually increasing in technical difficulty:
1. Synovial samples (liquid specimen, moderate to high concomitant debris such as hyaluronic acids, pus, and blood).
2. Biofilms on explants (solid specimen, low concomitant debris of tissue remnants and blood).
3. Tissue samples (solid specimen, high concomitant debris from solid tissue).
-
Transcriptome and microbiome
Diagnosis of orthopedic-related infections at the intersection of the transcriptome and microbiome.
Samir Mehta, University of Pennsylvania, United States
Osteomyelitis is a disease of heterogeneous presentation, etiology, management, and outcome which is among the most challenging complications in orthopedic surgery. In orthopedic trauma, the population most at risk of developing osteomyelitis includes open-fracture patients. Multiple microbial species are associated with either chronic or acute osteomyelitis (or both) but as these determinations typically depend on traditional culturing techniques, the potentially more complex etiology of osteomyelitis remains uncharacterized. Increased understanding of the specific microbial-host interaction between Staphylococcus aureus and human osteoblasts heightens the need for a refreshed approach to diagnose this disease in both the acute and chronic phases.
The overarching objective of the study is to use a system genomics approach to integrate microbiome and host transcriptome datasets to identify important biomarkers and develop predictive models of bone infection. The central hypothesis is that bone infections are associated with distinct microbial colonization and systemic host responses and analytical approaches that integrate these two variables together will provide more accurate and thorough interrogation of outcome associations. This hypothesis is based on studies demonstrating a previously unappreciated diversity of microbiota colonizing traumatic wounds and clear changes in microbial community structure, diversity, and quantity that precedes or coincides with infectious complications. Additionally, we have identified a preliminary panel of differentially expressed mRNA and miRNA from peripheral circulation that correlate with fracture outcome (healing vs nonunion). Included in this panel are a subset of mRNA transcripts that are representative of immune and inflammatory response.
We plan to test our central hypothesis with the following specific aims:
1. Identify features of the microbiome and host transcriptome that predict bone infection using a multi-omic integrative framework of host-microbe interactions.
2. Establish a predictive model of ongoing bone infection based on noninvasive profiling of host transcriptome signatures.
-
Immunological signatures for diagnosis of S. aureus bone infection
Evaluation of the immune proteome against Staphylococcus aureus (S. aureus) in patients with infected total joint replacements and development of a multiplex clinical immunoassay to assess the ongoing orthopedic infections and determine the probability of reinfection.
John L Daiss, Gowrishankar Muthukrishnan, University of Rochester Medical Center, United States
The collaboration between the leading institutes (Virginia Commonwealth University and University of Rochester) involved in the CPP has brought to the development of a multiplex immunoassay, to distinguish infected patients from healthy controls, which could be used as a diagnostic method. Described in detail in Nishitani et al (2015), such multiplex immunoassay is about 90 percent accurate in identifying S. aureus-infected patients, despite substantial variation among patients in terms of duration of infection and length and type of therapeutic intervention.
Every human adult has high levels of immunoglobulin G (IgG) specific for most of S. aureus antigens. This presumably reflects the immunological history of the subject who has experienced multiple overt or subclinical infections caused by S. aureus throughout his lifetime. In the multiplex immunoassay, background antibody levels that varied over two orders of magnitude were observed among healthy controls. Looking for a way to eliminate this background, the immunoassay was utilized to diagnose S. aureus infections with a novel analytic fluid.
Serum antibodies are the result of the patient’s immunological history and are produced by long-lived plasma cells in the bone marrow and spleen. In contrast, newly activated antibody secreting cells (ASC) emerge from the lymph nodes into the bloodstream only during ongoing infections and secrete pathogen-specific antibodies. By harvesting ASCs from whole blood and placing them in culture for several hours to enable them to secrete their antibodies, a novel analytic fluid, medium enriched for newly synthesized antibodies (MENSA), was created. Initial experiments with this novel analytic approach suggest that this assay can diagnose acute infections with 90 percent accuracy or more. This approach enables us to measure a patient’s long-term immune response to S. aureus infection in the serum and the immediate response to the ongoing infection in the MENSA. Therefore, we can explore both the fundamental nature of the immune response as well as its practical role as a diagnostic tool for orthopedic indications.
There are several new lines of experimental inquiry currently under exploration:
1. Are orthopedic infections different from skin or lung infections? Do they elicit a distinctive repertoire of cytokines or antibodies depending on the anatomical site of infection? Similarly, do infections on orthopedic implants elicit immune responses different from those in native bone?
2. What information about the infection is conveyed by the heavy chain class of the newly synthesized antibodies?
3. Do patients whose serum or MENSA antibody levels remain elevated throughout the months of treatment and recovery suffer worse outcomes than those who have experienced significant declines in antibody levels?
4. Do responses to particular antigens indicate success or failure of treatments? Alternatively, do responses to all antigens convey the same information?
5. Can MENSA antibody levels be used to monitor success or failure of therapy?
Other opportunities are also available for exploring clinical utility in complex and often difficult to sample orthopedic infections. For example, diabetic foot infections are a serious problem in the clinic. It is difficult to diagnose the identity of causative microorganisms due to the multiple species typically present in the wound. Success or failure of foot salvage therapy can be hard to track by external observations alone. Feasibility of MENSA in diagnosing diabetic foot infections and tracking foot salvage therapy in these patients was described by Oh et al (2018). Significantly, this line of inquiry can be extended to other species frequently involved in orthopedic infections such as S. epidermidis, S. lugdunensis, group B. streptococcus, Enterococcus, and P. aeruginosa.
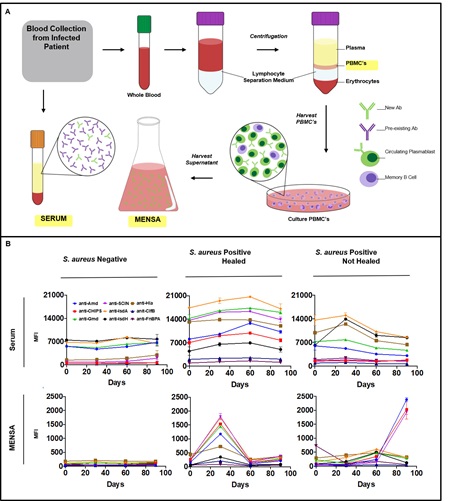
Figure: A diagnostic and prognostic immunoassay for the measurement of anti-S. aureus antibody levels in patients with osteomyelitis. (A) Schematic illustration of production of serum and isolation of MENSA from peripheral mononuclear cells of patients with osteomyelitis. (B) Anti-S. aureus antibody levels in serum and MENSA were determined using a custom bead-based multiantigen Luminex immunoassay developed by our group. Anti-S. aureus IgG responses was examined in serum and MENSA of diabetic foot infections (DFI) patients undergoing foot salvage antimicrobial therapy (FST). The change in antibody titers over the course of FST of a representative patient whose DFI was negative for S. aureus, a patient with S. aureus infection that responded to FST, and a patient with S. aureus DFI that failed FST are presented. Remarkably, MENSA levels faithfully reflected the S. aureus infection over time while serum levels remained unchanged.
The pillars of the CPP Bone Infection
Host-pathogen interaction
Learn about how S. aureus adapts to the environment of an implant-associated infection.
Diagnosis
Read about the development of new tools to assess implant-associated infections.
Prevention
Explore how the CPP Bone Infection is running its surgical site prevention program.
Publications
Check the publications of the CPP Bone Infection